Biopolymers 1.* (1995 F 6A) In biological systems adenosine triphosphate (ATP) serves as the principal immediate donor of free energy to make needed processes take place. It accomplishes this task by its hydrolysis in water to yield adenosine diphosphate (ADP) and orthophoshate (Pi) plus a proton (H+): ATP + H2O à
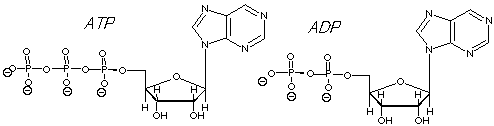
ADP + Pi + H+ For your information the structures of ATP and ADP are shown below:
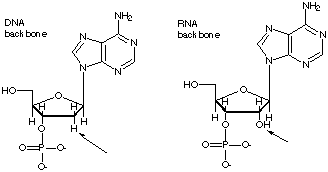
What features of ATP and ADP are also found in DNA and RNA? •The adenine base (a purine) in ATP is in one of the nuceotides in both DNA and RNA. •The five carbon ribose sugar in ATP is identical to that in RNA, while in DNA it is 2-deoxyribose (there is no hydroxyl group at the -2 carbon.) •Phosphodiesters, which link the sugar backbone in DNA and RNA, have the triphosphate group on ATP as their precursors.
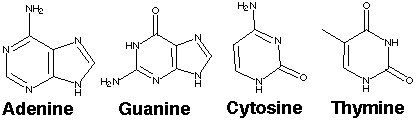
2.* (1995 F 8) The following are the four bases found in DNA:
B. The double-stranded helical structure of DNA is maintained primarily by the hydrogen bonds, which are weak bonds. With increasing heat, the double-stranded DNA can separate into single strands in a process called denaturation or "melting." The melting temperature Tm is defined as the temperature at which half the helical structure is lost. The abruptness of the transition indicates that the DNA double helix is a highly cooperative structure held together by many reinforcing bonds. Predict how the melting temperature varies with the base-pair composition in DNA for a given number of bases. G-C base pairs have 3 hydrogen bonds, while A-T base pairs have two. Therefore, double-stranded DNA with a higher number of G-C base pairs will be more strongly bonded together, more stable, and will have a higher melting temperature. 3.* (1995 2 2) Genes express proteins by transcription followed by translation.
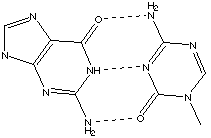
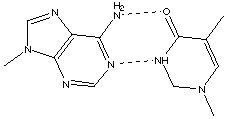
Transcription is the process by which the nucleotide sequence in a gene of DNA is copied into an RNA chain. RNA is synthesized by complementary base pairing with the template strand of DNA from the 5' end to the 3' end. DNA RNA A U T A C G G C Transcription starts with unwinding the DNA at the promoter region (beginning of the gene.) Translation begins by reading messenger RNA in the 5' to 3' direction according to the genetic code in which a triplet of bases (a codon) is assigned to one of the 20 amino acids. There is also a start codon and a stop codon. The start signal and the code for methionine are identical so that each protein begins with the amino acid methionine. There are 43=64 different possible codons so that more than one codon may code for the same amino acid, but a particular codon never specifies more than one amino acid. C. Often it is said that the change of one base for another in a gene causes a different protein to be expressed. Actually, more possibilities exist. Suggest how one base substitution might lead to (1) no change in the protein expressed, or (2) the expression of mor than one protein, or (3) the expression of no protein. • (1) no change in the protein expressed. There are redundant codons for many of the amino acids. Thus if one base is changed, it is possible that the affected codon will still code the same amino acid–thus the protein will not change. • (2) the expression of more than one protein. If a new start codon is created, then it may not be clear to the translation process which start codon to begin with. Thus one gene could create multiple proteins. • (3) the expression of no protein. If the start codon is changed to a non-start codon, or a stop codon is created right after the start codon, no protein will be formed. 4.* (1994 3 5) The four bases of the biopolymer DNA are A = adenine, T = thymine, G = guanine, and C = cytoseine. They form pairs, as shown below:
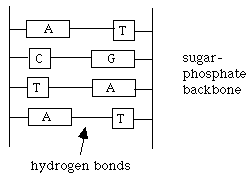
Draw a small portion of the DNA structure and explain these observations in terms of your schematic drawing.
DNA is a double helix, formed from two complementary strands. The sugar-phosphate backbone is on the outside, with the base pairs on the inside. Because of the hydrogen bonding, A always pairs with T, T always pairs with A, C always pairs with G, G always pairs with C, resulting in Chargaff's first two rules. Different DNA sequences lead to different mole fractions of A or G for various DNA's. B. When DNA is exposed to water at temperatures near its boiling point, this biopolymer falls apart in a process called denaturation. Experiments have shown that DNA containing more G (and C) than A (and T) are more stable, that is, they denature at higher temperatures. The denaturing temperature Td can be defined as the temperature at which one half the DNA is decomposed. Offer an explanation for the change in value of Td with G content. From the base-pairing diagram, we can see that the G-C pair has 3 hydrogen bonds, while the A-T pair has only 2. Therefore, the G-C pairing is more stable than the A-T pairing. Thus, strands with more G-C content have more hydrogen bonding, are more stable, and have a greater resistance to denaturation. Therefore, they require a higher Td to decompose. |